`moderndive`: Datasets and functions for `tidyverse`-friendly introductory linear regression
Albert Y. Kim, Chester Ismay, and Max Kuhn
2025-11-18
Source:vignettes/moderndive.Rmd
moderndive.RmdSummary
We present the moderndive R package of datasets and
functions for tidyverse-friendly introductory
linear regression (Wickham, Averick, et al.
2019). These tools leverage the well-developed
tidyverse and broom packages to facilitate 1)
working with regression tables that include confidence intervals, 2)
accessing regression outputs on an observation level
(e.g. fitted/predicted values and residuals), 3) inspecting scalar
summaries of regression fit
(e.g. ,
,
and mean squared error), and 4) visualizing parallel slopes regression
models using ggplot2-like syntax (Wickham, Chang, et al. 2019; Robinson and Hayes
2019). This R package is designed to supplement the book
“Statistical Inference via Data Science: A ModernDive into R and the
Tidyverse” (Ismay and Kim 2019). Note that
the book is also available online at https://moderndive.com and is referred to as
“ModernDive” for short.
Statement of Need
Linear regression has long been a staple of introductory statistics
courses. While the curricula of introductory statistics courses has much
evolved of late, the overall importance of regression remains the same
(American Statistical Association Undergraduate
Guidelines Workgroup 2016). Furthermore, while the use of the R
statistical programming language for statistical analysis is not new,
recent developments such as the tidyverse suite of packages
have made statistical computation with R accessible to a broader
audience (Wickham, Averick, et al. 2019).
We go one step further by leveraging the tidyverse and the
broom packages to make linear regression accessible to
students taking an introductory statistics course (Robinson and Hayes 2019). Such students are
likely to be new to statistical computation with R; we designed
moderndive with these students in mind.
Introduction
Let’s load all the R packages we are going to need.
Let’s consider data gathered from end of semester student evaluations
for a sample of 463 courses taught by 94 professors from the University
of Texas at Austin (Diez, Barr, and
Çetinkaya-Rundel 2015). This data is included in the
evals data frame from the moderndive
package.
In the following table, we present a subset of 9 of the 14 variables included for a random sample of 5 courses1:
-
IDuniquely identifies the course whereasprof_IDidentifies the professor who taught this course. This distinction is important since many professors taught more than one course. -
scoreis the outcome variable of interest: average professor evaluation score out of 5 as given by the students in this course. - The remaining variables are demographic variables describing that
course’s instructor, including
bty_avg(average “beauty” score) for that professor as given by a panel of 6 students.2
| ID | prof_ID | score | age | bty_avg | gender | ethnicity | language | rank |
|---|---|---|---|---|---|---|---|---|
| 129 | 23 | 3.7 | 62 | 3.000 | male | not minority | english | tenured |
| 109 | 19 | 4.7 | 46 | 4.333 | female | not minority | english | tenured |
| 28 | 6 | 4.8 | 62 | 5.500 | male | not minority | english | tenured |
| 434 | 88 | 2.8 | 62 | 2.000 | male | not minority | english | tenured |
| 330 | 66 | 4.0 | 64 | 2.333 | male | not minority | english | tenured |
Regression analysis the “good old-fashioned” way
Let’s fit a simple linear regression model of teaching
score as a function of instructor age using
the lm() function.
score_model <- lm(score ~ age, data = evals)Let’s now study the output of the fitted model
score_model “the good old-fashioned way”: using
summary() which calls summary.lm() behind the
scenes (we’ll refer to them interchangeably throughout this paper).
summary(score_model)
##
## Call:
## lm(formula = score ~ age, data = evals)
##
## Residuals:
## Min 1Q Median 3Q Max
## -1.9185 -0.3531 0.1172 0.4172 0.8825
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 4.461932 0.126778 35.195 <2e-16 ***
## age -0.005938 0.002569 -2.311 0.0213 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.5413 on 461 degrees of freedom
## Multiple R-squared: 0.01146, Adjusted R-squared: 0.009311
## F-statistic: 5.342 on 1 and 461 DF, p-value: 0.02125Here are 5 common student questions we’ve heard over the years in our introductory statistics courses based on this output:
- “Wow! Look at those p-value stars! Stars are good, so I should try to get many stars, right?”
- “How do we extract the values in the regression table?”
- “Where are the fitted/predicted values and residuals?”
- “How do I apply this model to a new set of data to make predictions?”
- “What is all this other stuff at the bottom?”
Regression analysis using moderndive
To address these questions, we’ve included three functions in the
moderndive package that take a fitted model object as input
and return the same information as summary.lm(), but output
them in tidyverse-friendly format (Wickham,
Averick, et al. 2019). As we’ll see later, while these three
functions are thin wrappers to existing functions in the
broom package for converting statistical objects into tidy
tibbles, we modified them with the introductory statistics student in
mind (Robinson and Hayes 2019).
-
Get a tidy regression table with confidence intervals:
get_regression_table(score_model) ## # A tibble: 2 × 7 ## term estimate std_error statistic p_value lower_ci upper_ci ## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> ## 1 intercept 4.46 0.127 35.2 0 4.21 4.71 ## 2 age -0.006 0.003 -2.31 0.021 -0.011 -0.001 -
Get information on each point/observation in your regression, including fitted/predicted values and residuals, in a single data frame:
get_regression_points(score_model) ## # A tibble: 463 × 5 ## ID score age score_hat residual ## <int> <dbl> <int> <dbl> <dbl> ## 1 1 4.7 36 4.25 0.452 ## 2 2 4.1 36 4.25 -0.148 ## 3 3 3.9 36 4.25 -0.348 ## 4 4 4.8 36 4.25 0.552 ## 5 5 4.6 59 4.11 0.488 ## 6 6 4.3 59 4.11 0.188 ## 7 7 2.8 59 4.11 -1.31 ## 8 8 4.1 51 4.16 -0.059 ## 9 9 3.4 51 4.16 -0.759 ## 10 10 4.5 40 4.22 0.276 ## # ℹ 453 more rows -
Get scalar summaries of a regression fit including and but also the (root) mean-squared error:
get_regression_summaries(score_model) ## # A tibble: 1 × 9 ## r_squared adj_r_squared mse rmse sigma statistic p_value df ## <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> ## 1 0.011 0.009 0.292 0.540 0.541 5.34 0.021 1 ## # ℹ 1 more variable: nobs <dbl>
Bonus: Visualizing parallel slopes models with
moderndive
Furthermore, say you would like to create a visualization of the
relationship between two numerical variables and a third categorical
variable with
levels. Let’s create this using a colored scatterplot via the
ggplot2 package for data visualization (Wickham, Chang, et al. 2019). Using
geom_smooth(method = "lm", se = FALSE) yields a
visualization of an interaction model where each of the
regression lines has their own intercept and slope. For example in , we
extend our previous regression model by now mapping the categorical
variable ethnicity to the color aesthetic.
# Code to visualize interaction model:
ggplot(evals, aes(x = age, y = score, color = ethnicity)) +
geom_point() +
geom_smooth(method = "lm", se = FALSE) +
labs(x = "Age", y = "Teaching score", color = "Ethnicity")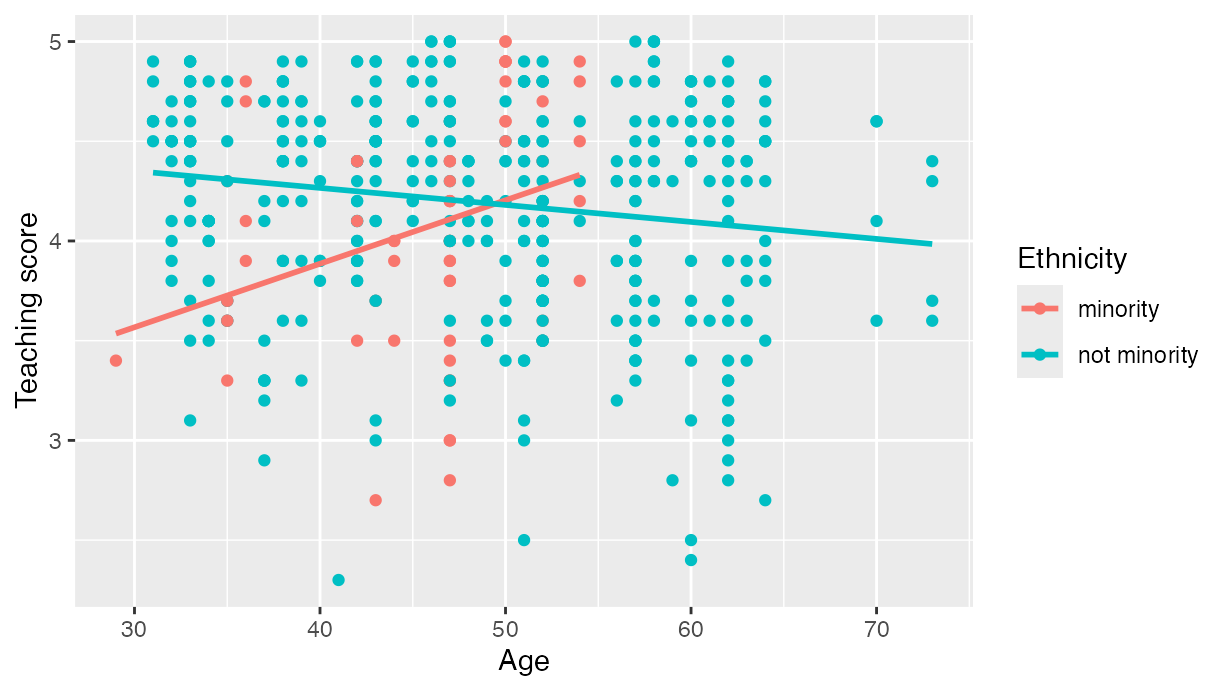
Visualization of interaction model.
However, many introductory statistics courses start with the easier
to teach “common slope, different intercepts” regression model, also
known as the parallel slopes model. However, no argument to
plot such models exists within geom_smooth().
Evgeni
Chasnovski thus wrote a custom geom_ extension to
ggplot2 called geom_parallel_slopes(); this
extension is included in the moderndive package. Much like
geom_smooth() from the ggplot2 package, you
add geom_parallel_slopes() as a layer to the code,
resulting in .
# Code to visualize parallel slopes model:
ggplot(evals, aes(x = age, y = score, color = ethnicity)) +
geom_point() +
geom_parallel_slopes(se = FALSE) +
labs(x = "Age", y = "Teaching score", color = "Ethnicity")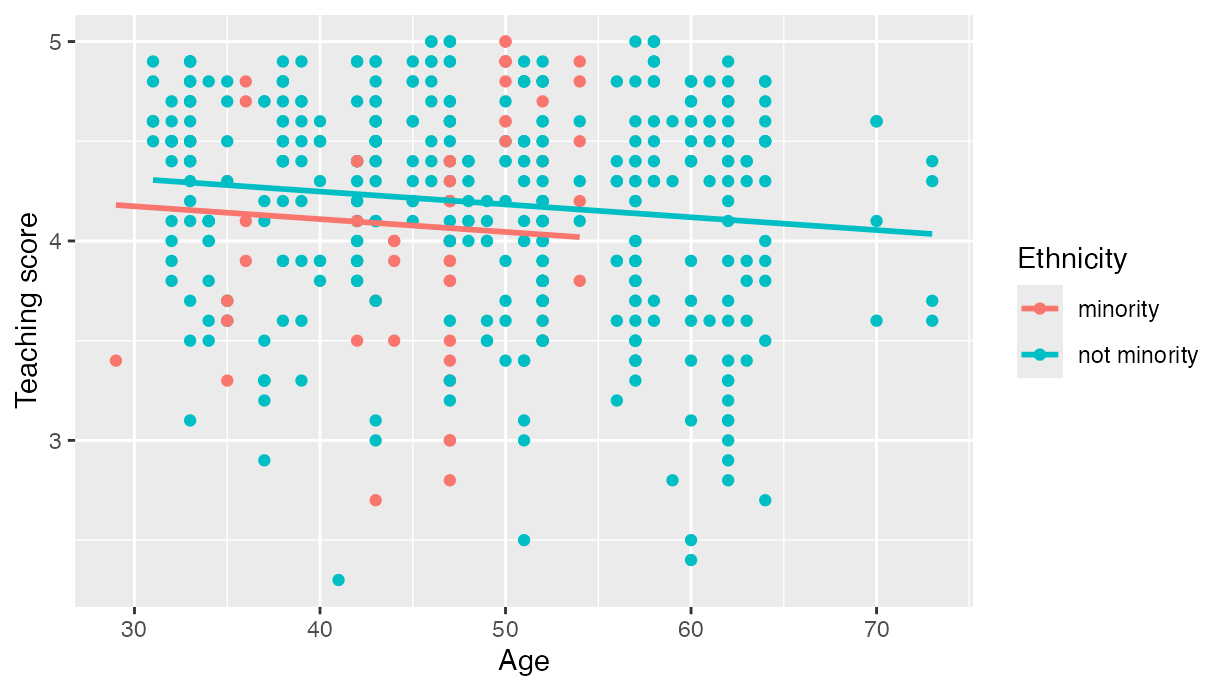
Visualization of parallel slopes model.
At this point however, students will inevitably ask a sixth question: “When would you ever use a parallel slopes model?”
Why should you use the moderndive package?
To recap this introduction, we believe that the following functions
included in the moderndive package
are effective pedagogical tools that can help address the six common questions posed by students about introductory linear regression performed in R. We now argue why.
Features
1. Less p-value stars, more confidence intervals
The first common student question:
“Wow! Look at those p-value stars! Stars are good, so I should try to get many stars, right?”
We argue that the summary.lm() output is deficient in an
introductory statistics setting because:
- The
Signif. codes: 0 '' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1only encourage p-hacking. In case you have not yet been convinced of the perniciousness of p-hacking, perhaps comedian John Oliver can convince you.
- While not a silver bullet for eliminating misinterpretations of
statistical inference, confidence intervals present students with a
sense of the associated effect sizes of any explanatory variables. Thus,
practical as well as statistical significance is emphasized. These are
not included by default in the output of
summary.lm().
Instead of summary(), let’s use the
get_regression_table() function:
get_regression_table(score_model)
## # A tibble: 2 × 7
## term estimate std_error statistic p_value lower_ci upper_ci
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 intercept 4.46 0.127 35.2 0 4.21 4.71
## 2 age -0.006 0.003 -2.31 0.021 -0.011 -0.001Observe how the p-value stars are omitted and confidence intervals
for the point estimates of all regression parameters are included by
default. By including them in the output, we can then emphasize to
students that they “surround” the point estimates in the
estimate column. Note the confidence level is defaulted to
95%; this default can be changed using the conf.level
argument:
get_regression_table(score_model, conf.level = 0.99)
## # A tibble: 2 × 7
## term estimate std_error statistic p_value lower_ci upper_ci
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 intercept 4.46 0.127 35.2 0 4.13 4.79
## 2 age -0.006 0.003 -2.31 0.021 -0.013 0.0012. Outputs as tibbles
The second common student question:
“How do we extract the values in the regression table?”
While one might argue that extracting the intercept and slope
coefficients can be “simply” done using
coefficients(score_model), what about the standard errors?
For example, a Google query of “how do I extract standard errors
from lm in R” yielded results from the R mailing list and from Cross Validated suggesting we run:
We argue that this task shouldn’t be this hard, especially in an
introductory statistics setting. To rectify this, the three
get_regression_* functions all return data frames in the
tidyverse-style tibble (tidy table) format (Müller and Wickham 2019). Therefore you can
extract columns using the pull() function from the
dplyr package (Wickham et al.
2020):
get_regression_table(score_model) %>%
pull(std_error)
## [1] 0.127 0.003or equivalently you can use the $ sign operator from
base R:
get_regression_table(score_model)$std_error
## [1] 0.127 0.003Furthermore, by piping the above
get_regression_table(score_model) output into the
kable() function from the knitr package (Xie 2020), you can obtain aesthetically
pleasing regression tables in R Markdown documents, instead of tables
written in jarring computer output font:
get_regression_table(score_model) %>%
kable()| term | estimate | std_error | statistic | p_value | lower_ci | upper_ci |
|---|---|---|---|---|---|---|
| intercept | 4.462 | 0.127 | 35.195 | 0.000 | 4.213 | 4.711 |
| age | -0.006 | 0.003 | -2.311 | 0.021 | -0.011 | -0.001 |
3. Birds of a feather should flock together: Fitted values & residuals
The third common student question:
“Where are the fitted/predicted values and residuals?”
How can we extract point-by-point information from a regression model, such as the fitted/predicted values and the residuals? (Note we only display the first 10 out of 463 of such values for brevity’s sake.)
fitted(score_model)## 1 2 3 4 5 6 7
## 4.248156 4.248156 4.248156 4.248156 4.111577 4.111577 4.111577
## 8 9 10
## 4.159083 4.159083 4.224403
residuals(score_model)## 1 2 3 4 5
## 0.45184376 -0.14815624 -0.34815624 0.55184376 0.48842294
## 6 7 8 9 10
## 0.18842294 -1.31157706 -0.05908286 -0.75908286 0.27559666But why have the original explanatory/predictor age and
outcome variable score in evals, the
fitted/predicted values score_hat, and
residual floating around in separate vectors? Since each
observation relates to the same course, we argue it makes more sense to
organize them together in the same data frame using
get_regression_points():
score_model_points <- get_regression_points(score_model)
score_model_points## # A tibble: 10 × 5
## ID score age score_hat residual
## <int> <dbl> <int> <dbl> <dbl>
## 1 1 4.7 36 4.25 0.452
## 2 2 4.1 36 4.25 -0.148
## 3 3 3.9 36 4.25 -0.348
## 4 4 4.8 36 4.25 0.552
## 5 5 4.6 59 4.11 0.488
## 6 6 4.3 59 4.11 0.188
## 7 7 2.8 59 4.11 -1.31
## 8 8 4.1 51 4.16 -0.059
## 9 9 3.4 51 4.16 -0.759
## 10 10 4.5 40 4.22 0.276Observe that the original outcome variable score and
explanatory/predictor variable age are now supplemented
with the fitted/predicted values score_hat and
residual columns. By putting the fitted values, predicted
values, and residuals next to the original data, we argue that the
computation of these values is less opaque. For example, instructors can
emphasize how all values in the first row of output are computed.
Furthermore, recall that since all outputs in the
moderndive package are tibble data frames, custom residual
analysis plots can be created instead of relying on the default plots
yielded by plot.lm(). For example, we can check for the
normality of residuals using the histogram of residuals shown in .
# Code to visualize distribution of residuals:
ggplot(score_model_points, aes(x = residual)) +
geom_histogram(bins = 20) +
labs(x = "Residual", y = "Count")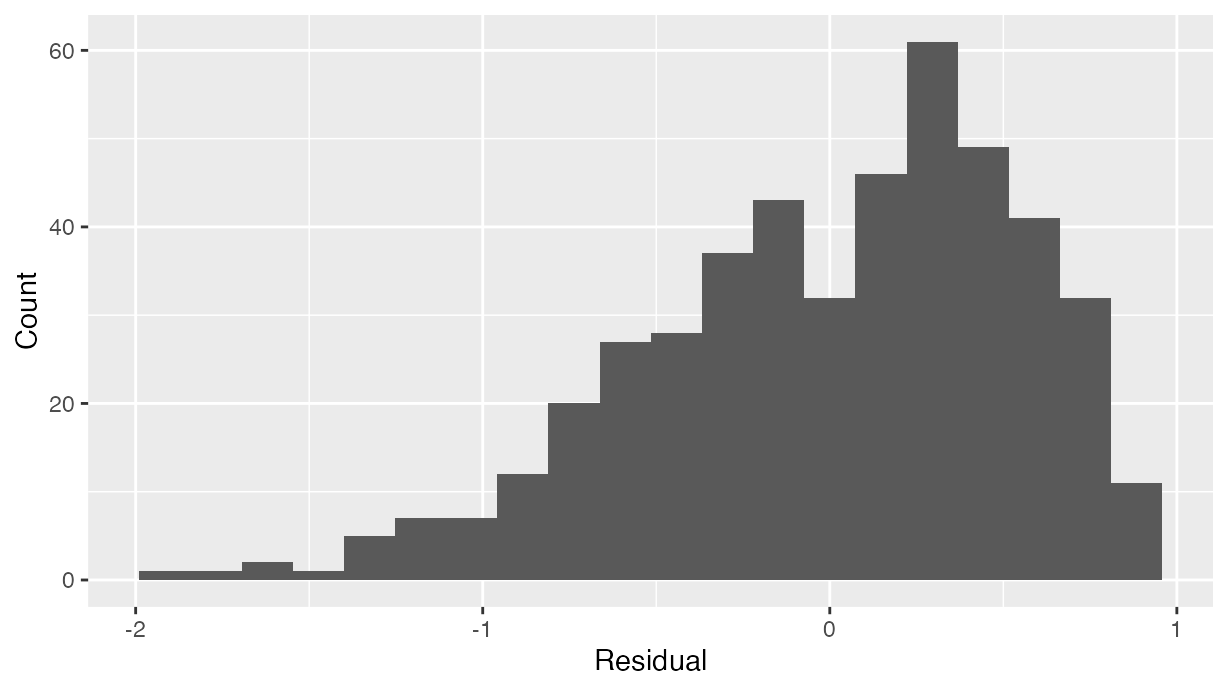
Histogram visualizing distribution of residuals.
As another example, we can investigate potential relationships between the residuals and all explanatory/predictor variables and the presence of heteroskedasticity using partial residual plots, like the partial residual plot over age shown in . If the term “heteroskedasticity” is new to you, it corresponds to the variability of one variable being unequal across the range of values of another variable. The presence of heteroskedasticity violates one of the assumptions of inference for linear regression.
# Code to visualize partial residual plot over age:
ggplot(score_model_points, aes(x = age, y = residual)) +
geom_point() +
labs(x = "Age", y = "Residual")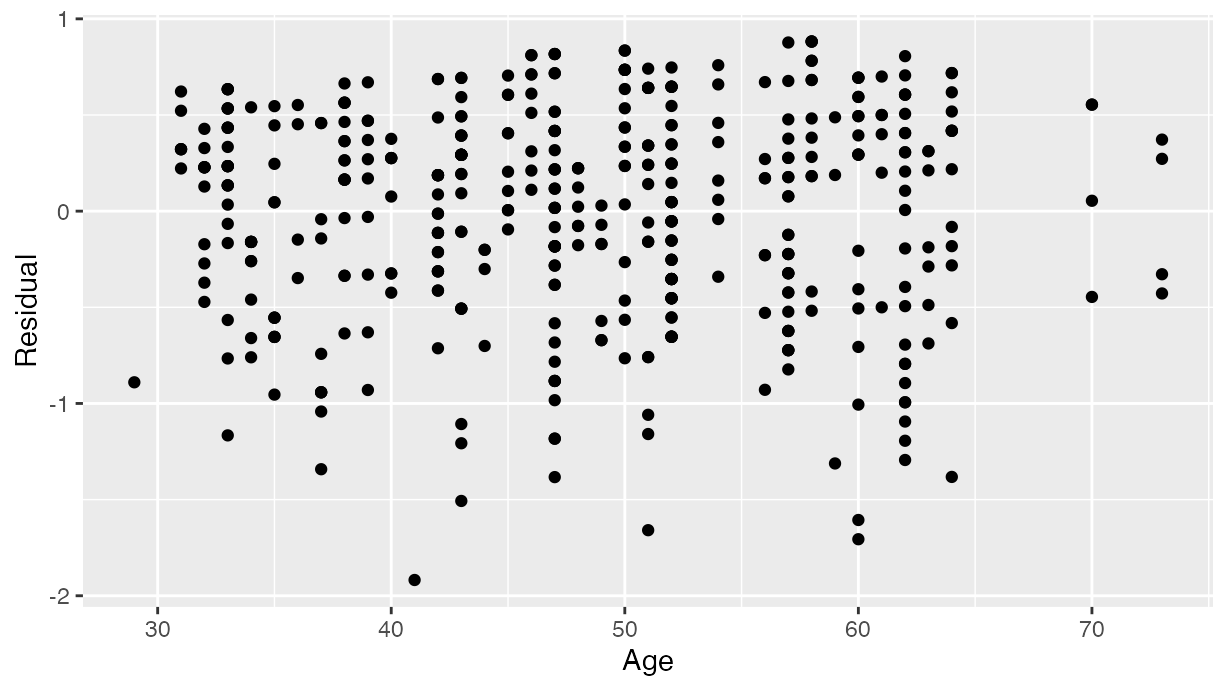
Partial residual residual plot over age.
4. A quick-and-easy Kaggle predictive modeling competition submission!
The fourth common student question:
“How do I apply this model to a new set of data to make predictions?”
With the fields of machine learning and artificial intelligence
gaining prominence, the importance of predictive modeling cannot be
understated. Therefore, we’ve designed the
get_regression_points() function to allow for a
newdata argument to quickly apply a previously fitted model
to new observations.
Let’s create an artificial “new” dataset consisting of two
instructors of age 39 and 42 and save it in a tibble data frame called
new_prof. We then set the newdata argument to
get_regression_points() to apply our previously fitted
model score_model to this new data, where
score_hat holds the corresponding fitted/predicted
values.
new_prof <- tibble(age = c(39, 42))
get_regression_points(score_model, newdata = new_prof)
## # A tibble: 2 × 3
## ID age score_hat
## <int> <dbl> <dbl>
## 1 1 39 4.23
## 2 2 42 4.21Let’s do another example, this time using the Kaggle House Prices: Advanced Regression Techniques practice competition ( displays the homepage for this competition).
House prices Kaggle competition homepage.
This Kaggle competition requires you to fit/train a model to the
provided train.csv training set to make predictions of
house prices in the provided test.csv test set. We present
an application of the get_regression_points() function
allowing students to participate in this Kaggle competition. It
will:
- Read in the training and test data.
- Fit a naive model of house sale price as a function of year sold to the training data.
- Make predictions on the test data and write them to a
submission.csvfile that can be submitted to Kaggle usingget_regression_points(). Note the use of theIDargument to use theidvariable intestto identify the rows (a requirement of Kaggle competition submissions).
library(readr)
library(dplyr)
library(moderndive)
# Load in training and test set
train <- read_csv("https://moderndive.com/data/train.csv")
test <- read_csv("https://moderndive.com/data/test.csv")
# Fit model:
house_model <- lm(SalePrice ~ YrSold, data = train)
# Make predictions and save in appropriate data frame format:
submission <- house_model %>%
get_regression_points(newdata = test, ID = "Id") %>%
select(Id, SalePrice = SalePrice_hat)
# Write predictions to csv:
write_csv(submission, "submission.csv")After submitting submission.csv to the leaderboard for
this Kaggle competition, we obtain a “root mean squared logarithmic
error” (RMSLE) score of 0.42918 as seen in .
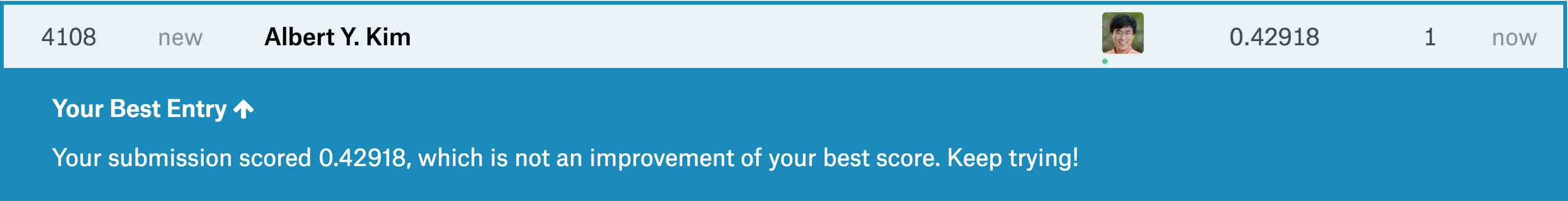
Resulting Kaggle RMSLE score.
5. Scalar summaries of linear regression model fits
The fifth common student question:
“What is all this other stuff at the bottom?”
Recall the output of the standard summary.lm() from
earlier:
##
## Call:
## lm(formula = score ~ age, data = evals)
##
## Residuals:
## Min 1Q Median 3Q Max
## -1.9185 -0.3531 0.1172 0.4172 0.8825
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 4.461932 0.126778 35.195 <2e-16 ***
## age -0.005938 0.002569 -2.311 0.0213 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.5413 on 461 degrees of freedom
## Multiple R-squared: 0.01146, Adjusted R-squared: 0.009311
## F-statistic: 5.342 on 1 and 461 DF, p-value: 0.02125Say we wanted to extract the scalar model summaries at the bottom of
this output, such as
,
,
and the
-statistic.
We can do so using the get_regression_summaries()
function.
get_regression_summaries(score_model)
## # A tibble: 1 × 9
## r_squared adj_r_squared mse rmse sigma statistic p_value df
## <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 0.011 0.009 0.292 0.540 0.541 5.34 0.021 1
## # ℹ 1 more variable: nobs <dbl>We’ve supplemented the standard scalar summaries output yielded by
summary() with the mean squared error mse and
root mean squared error rmse given their popularity in
machine learning settings.
6. Plot parallel slopes regression models
Finally, the last common student question:
“When would you ever use a parallel slopes model?”
For example, recall the earlier visualizations of the interaction and parallel slopes models for teaching score as a function of age and ethnicity we saw in Figures and . Let’s present both visualizations side-by-side in .
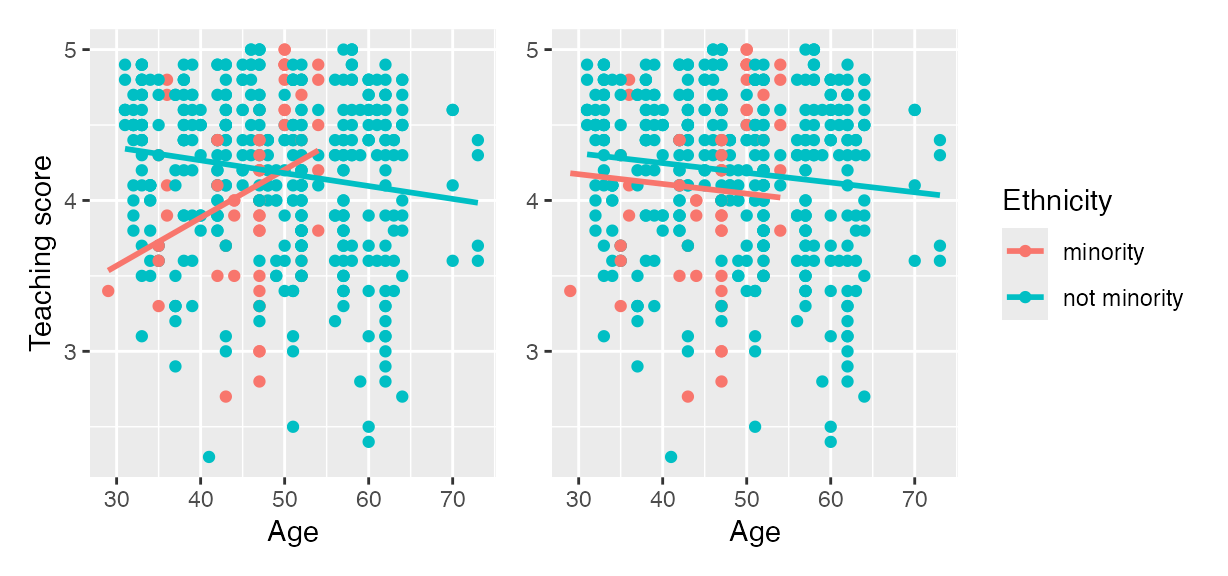
Interaction (left) and parallel slopes (right) models.
Students might be wondering “Why would you use the parallel slopes model on the right when the data clearly form an”X” pattern as seen in the interaction model on the left?” This is an excellent opportunity to gently introduce the notion of model selection and Occam’s Razor: an interaction model should only be used over a parallel slopes model if the additional complexity of the interaction model is warranted. Here, we define model “complexity/simplicity” in terms of the number of parameters in the corresponding regression tables:
# Regression table for interaction model:
interaction_evals <- lm(score ~ age * ethnicity, data = evals)
get_regression_table(interaction_evals)
## # A tibble: 4 × 7
## term estimate std_error statistic p_value lower_ci upper_ci
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 intercept 2.61 0.518 5.04 0 1.59 3.63
## 2 age 0.032 0.011 2.84 0.005 0.01 0.054
## 3 ethnicity-no… 2.00 0.534 3.74 0 0.945 3.04
## 4 age:ethnicit… -0.04 0.012 -3.51 0 -0.063 -0.018
# Regression table for parallel slopes model:
parallel_slopes_evals <- lm(score ~ age + ethnicity, data = evals)
get_regression_table(parallel_slopes_evals)
## # A tibble: 3 × 7
## term estimate std_error statistic p_value lower_ci upper_ci
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 intercept 4.37 0.136 32.1 0 4.1 4.63
## 2 age -0.006 0.003 -2.5 0.013 -0.012 -0.001
## 3 ethnicity-no… 0.138 0.073 1.89 0.059 -0.005 0.282The interaction model is “more complex” as evidenced by its regression table involving 4 rows of parameter estimates whereas the parallel slopes model is “simpler” as evidenced by its regression table involving only 3 parameter estimates. It can be argued however that this additional complexity is warranted given the clearly different slopes in the left-hand plot of .
We now present a contrasting example, this time from Chapter 6 of the online version of ModernDive Subsection 6.3.1 involving Massachusetts USA public high schools.3 Let’s plot both the interaction and parallel slopes models in .
# Code to plot interaction and parallel slopes models for MA_schools
ggplot(
MA_schools,
aes(x = perc_disadvan, y = average_sat_math, color = size)
) +
geom_point(alpha = 0.25) +
labs(
x = "% economically disadvantaged",
y = "Math SAT Score",
color = "School size"
) +
geom_smooth(method = "lm", se = FALSE)
ggplot(
MA_schools,
aes(x = perc_disadvan, y = average_sat_math, color = size)
) +
geom_point(alpha = 0.25) +
labs(
x = "% economically disadvantaged",
y = "Math SAT Score",
color = "School size"
) +
geom_parallel_slopes(se = FALSE)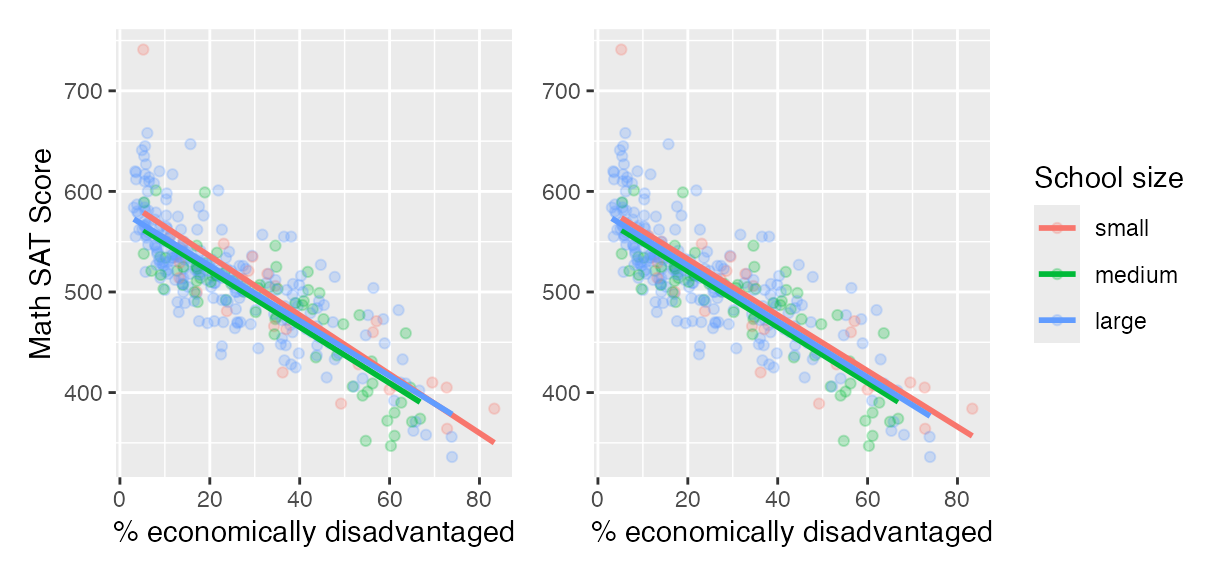
Interaction (left) and parallel slopes (right) models.
In terms of the corresponding regression tables, observe that the corresponding regression table for the parallel slopes model has 4 rows as opposed to the 6 for the interaction model, reflecting its higher degree of “model simplicity.”
# Regression table for interaction model:
interaction_MA <-
lm(average_sat_math ~ perc_disadvan * size, data = MA_schools)
get_regression_table(interaction_MA)
## # A tibble: 6 × 7
## term estimate std_error statistic p_value lower_ci upper_ci
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 intercept 594. 13.3 44.7 0 568. 620.
## 2 perc_disadvan -2.93 0.294 -9.96 0 -3.51 -2.35
## 3 size-medium -17.8 15.8 -1.12 0.263 -48.9 13.4
## 4 size-large -13.3 13.8 -0.962 0.337 -40.5 13.9
## 5 perc_disadva… 0.146 0.371 0.393 0.694 -0.585 0.877
## 6 perc_disadva… 0.189 0.323 0.586 0.559 -0.446 0.824
# Regression table for parallel slopes model:
parallel_slopes_MA <-
lm(average_sat_math ~ perc_disadvan + size, data = MA_schools)
get_regression_table(parallel_slopes_MA)
## # A tibble: 4 × 7
## term estimate std_error statistic p_value lower_ci upper_ci
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 intercept 588. 7.61 77.3 0 573. 603.
## 2 perc_disadvan -2.78 0.106 -26.1 0 -2.99 -2.57
## 3 size-medium -11.9 7.54 -1.58 0.115 -26.7 2.91
## 4 size-large -6.36 6.92 -0.919 0.359 -20.0 7.26Unlike our earlier comparison of interaction and parallel slopes models in , in this case it could be argued that the additional complexity of the interaction model is not warranted since the 3 three regression lines in the left-hand interaction are already somewhat parallel. Therefore the simpler parallel slopes model should be favored.
Going one step further, notice how the three regression lines in the
visualization of the parallel slopes model in the right-hand plot of
have similar intercepts. In can thus be argued that the additional model
complexity induced by introducing the categorical variable school
size is not warranted. Therefore, a simple linear
regression model using only perc_disadvan percent of the
student body that are economically disadvantaged should be favored.
While many students will inevitably find these results depressing, in our opinion, it is important to additionally emphasize that such regression analyses can be used as an empowering tool to bring to light inequities in access to education and inform policy decisions.
The Details
Three wrappers to broom functions
As we mentioned earlier, the three get_regression_*
functions are wrappers of functions from the broom package
for converting statistical analysis objects into tidy tibbles along with
a few added tweaks, but with the introductory statistics student in mind
(Robinson and Hayes 2019):
-
get_regression_table()is a wrapper forbroom::tidy(). -
get_regression_points()is a wrapper forbroom::augment(). -
get_regression_summariesis a wrapper forbroom::glance().
Why did we take this approach to address the initial 5 common student questions at the outset of the article?
- By writing wrappers to pre-existing functions, instead of creating new custom functions, there is minimal re-inventing of the wheel necessary.
- In our experience, novice R users had a hard time understanding the
broompackage function namestidy(),augment(), andglance(). To make them more user-friendly, themoderndivepackage wrappers have much more intuitively namedget_regression_table(),get_regression_points(), andget_regression_summaries(). - The variables included in the outputs of the above 3
broomfunctions are not all applicable to an introductory statistics students and of those that were, we found they were not intuitive for new R users. We therefore cut out some of the variables from the output and renamed some of the remaining variables. For example, compare the outputs of theget_regression_points()wrapper function and the parentbroom::augment()function.
get_regression_points(score_model)
## # A tibble: 463 × 5
## ID score age score_hat residual
## <int> <dbl> <int> <dbl> <dbl>
## 1 1 4.7 36 4.25 0.452
## 2 2 4.1 36 4.25 -0.148
## 3 3 3.9 36 4.25 -0.348
## 4 4 4.8 36 4.25 0.552
## 5 5 4.6 59 4.11 0.488
## 6 6 4.3 59 4.11 0.188
## 7 7 2.8 59 4.11 -1.31
## 8 8 4.1 51 4.16 -0.059
## 9 9 3.4 51 4.16 -0.759
## 10 10 4.5 40 4.22 0.276
## # ℹ 453 more rows
broom::augment(score_model)
## # A tibble: 463 × 8
## score age .fitted .resid .hat .sigma .cooksd .std.resid
## <dbl> <int> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 4.7 36 4.25 0.452 0.00560 0.542 0.00197 0.837
## 2 4.1 36 4.25 -0.148 0.00560 0.542 0.000212 -0.274
## 3 3.9 36 4.25 -0.348 0.00560 0.542 0.00117 -0.645
## 4 4.8 36 4.25 0.552 0.00560 0.541 0.00294 1.02
## 5 4.6 59 4.11 0.488 0.00471 0.541 0.00193 0.904
## 6 4.3 59 4.11 0.188 0.00471 0.542 0.000288 0.349
## 7 2.8 59 4.11 -1.31 0.00471 0.538 0.0139 -2.43
## 8 4.1 51 4.16 -0.0591 0.00232 0.542 0.0000139 -0.109
## 9 3.4 51 4.16 -0.759 0.00232 0.541 0.00229 -1.40
## 10 4.5 40 4.22 0.276 0.00374 0.542 0.000488 0.510
## # ℹ 453 more rowsThe source code for these three get_regression_*
functions can be found on GitHub.
Custom geometries
The geom_parallel_slopes() is a custom built
geom extension to the ggplot2 package. For
example, the ggplot2 webpage page gives instructions on how to create such extensions. The
source code for geom_parallel_slopes() written by Evgeni
Chasnovski can be found on GitHub.
Author contributions
Albert Y. Kim and Chester Ismay contributed equally to the
development of the moderndive package. Albert Y. Kim wrote
a majority of the initial version of this manuscript with Chester Ismay
writing the rest. Max Kuhn provided guidance and feedback at various
stages of the package development and manuscript writing.
Acknowledgments
Many thanks to Jenny Smetzer @smetzer180, Luke W. Johnston @lwjohnst86,
and Lisa Rosenthal @lisamr for their helpful feedback for this vignette
and to Evgeni Chasnovski @echasnovski for contributing the
geom_parallel_slopes() function via GitHub pull request. The authors do not have any financial
support to disclose.